Oilfield & Energies
Use of antibiotic combinations in fuel ethanol production
26th June 2019
By Jennifer Starner, PhD, Research Scientist, U.S. Water
Bacterial infections during fuel ethanol fermentations can negatively impact the fermentation process and lower ethanol yields.&nb
Bacterial infections during fuel ethanol fermentations can negatively impact the fermentation process and lower ethanol yields. Jennifer Starner, PhD Research Scientist at U.S. Water, explains how the use of a single antibiotic at a time with periodic switching to an alternative antibiotic, provides the most effective bacterial control while limiting the development of multi-drug resistance.
Lactic acid producing bacteria (LAB), such as Lactobacillus, Lactococcus, Weissella and Pediococcus, are a common source of contamination during fuel ethanol fermentations due to their ability to grow in the harsh conditions with low pH, high temperatures and high concentrations of ethanol. LAB are problematic as they not only compete with the yeast for nutrients and fermentable sugars, they also produce lactic and acetic acids that influence yeast metabolism. Ultimately, untreated bacterial infections cost plants time and money. Antibiotics, such as the β-lactam antibiotic penicillin G and the streptogramin antibiotic virginiamycin, are commonly used during fermentations to control bacterial growth and support robust ethanol yields.
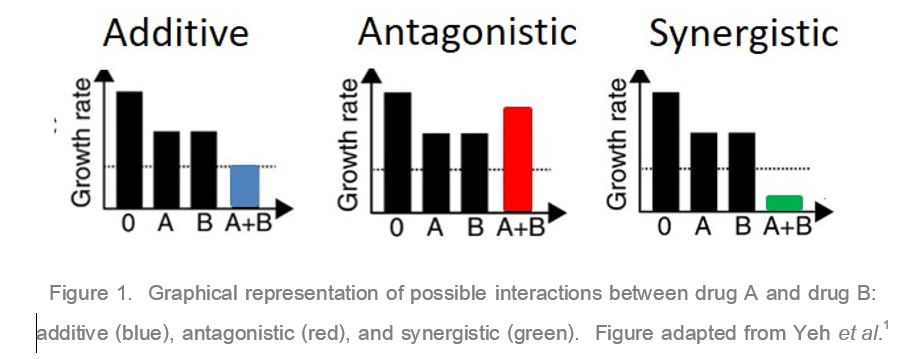 In response to a competitive antibiotic market and increasing antibiotic resistance, antibiotic combinations, particularly combinations of penicillin G and virginiamycin, have recently been employed. However, there is little evidence supporting the use of a penicillin G/virginiamycin combination treatment during fuel ethanol fermentations. When drugs such as antibiotics are applied concurrently, drug interactions must be considered. Drug interactions can be classified as additive, synergistic, or antagonistic (Figure 1). An additive interaction is when the two drugs do not interact and the response to the combination is the sum of the individual responses. Drugs are synergistic when application of the two drugs together creates a greater response than either drug individually. In direct contrast, drugs are antagonistic when the combination leads to a lesser response compared to the individual constituents.
In response to a competitive antibiotic market and increasing antibiotic resistance, antibiotic combinations, particularly combinations of penicillin G and virginiamycin, have recently been employed. However, there is little evidence supporting the use of a penicillin G/virginiamycin combination treatment during fuel ethanol fermentations. When drugs such as antibiotics are applied concurrently, drug interactions must be considered. Drug interactions can be classified as additive, synergistic, or antagonistic (Figure 1). An additive interaction is when the two drugs do not interact and the response to the combination is the sum of the individual responses. Drugs are synergistic when application of the two drugs together creates a greater response than either drug individually. In direct contrast, drugs are antagonistic when the combination leads to a lesser response compared to the individual constituents.Synergistic drug pairs are rare, occurring in only 4–10% of drug pairs tested, and if the drugs target different metabolic pathways the chance of antagonism increases.2 Penicillin and virginiamycin target very different metabolic pathways within the bacteria cell. Penicillin G inhibits cell wall synthesis, while virginiamycin inhibits protein synthesis by targeting the 50S ribosomal subunit. There is significant research examining how penicillin interacts with other antibiotics. Penicillin has a mixed response profile; it is synergistic with some classes of antibiotics such as aminoglycosides and antagonistic with others, particularly antagonistic with protein synthesis inhibitors that target the 50S ribosomal subunit, such as virginiamycin.1,3
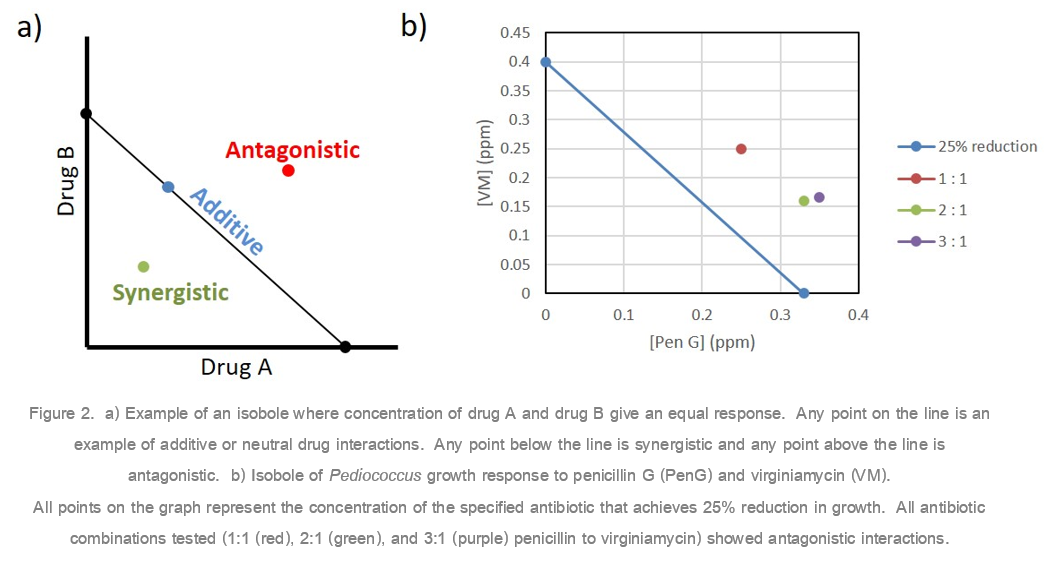
To explore the relationship between penicillin and virginiamycin when used as a combination product, we compared the efficacy of each individual antibiotic and various combinations against both lab bacterial strains and isolates collected from operating ethanol plants. Pediococcus pentosaceus, a common bacteria found in ethanol fermentations, was grown in the presence of either no antibiotic, penicillin G or virginiamycin alone, or 1:1, 2:1, or 3:1 ratios of penicillin to virginiamycin. Isobolograms, a common tool used to analyze drug combination results, were used to determine whether the drug combinations were synergistic (more effective), additive (neutral), or antagonistic (less effective).4 Isobolograms look at the concentration of the antibiotic required to achieve a desired response, such as 25% reduction in growth when compared to no antibiotic. If the drug combination is synergistic, the concentration of the antibiotics required to achieve that response will be lower than the individual antibiotic concentrations. If the drug combination is antagonistic, a greater concentration will be required to achieve that response (Figure 2a). Pediococcus pentosaceus results clearly showed that the penicillin/virginiamycin combinations, regardless of the ratio used, were antagonistic and on an equal dose comparison the individual antibiotics were more effective at controlling bacterial growth (Figure 2b).
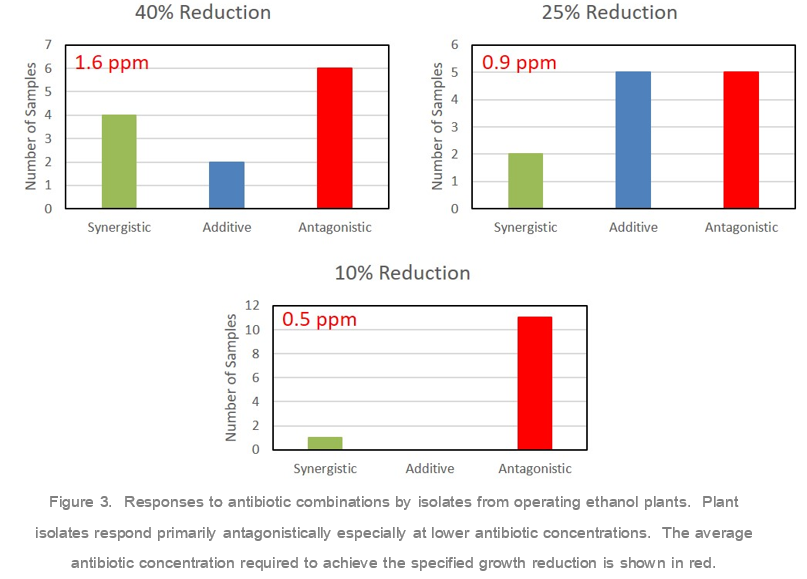
In addition to testing a lab strain, we also tested twelve isolates from eight different operating ethanol plants over the past year. As with the lab bacterial strain, isolates were grown in the presence of either no antibiotic, each individual antibiotic, or various combinations. For analysis of the results, we chose to focus on 40%, 25%, and 10% growth reduction responses as the average antibiotic concentration required correlated with the typical antibiotic doses used in the field, 1.6, 0.9, and 0.5 ppm respectively (Figure 3). While there were a few examples of synergy seen with the pencillin/virginiamycin combinations, the results overall showed a significant trend toward antagonism. This becomes especially true at lower antibiotic concentrations. When looking at the 10% growth reduction isoboles, which corresponds to 0.5 ppm average antibiotic dose, eleven of the twelve isolates reacted antagonistically with the combination products. Our results show that antibiotic combinations are less effective at controlling bacterial growth compared to the individual antibiotics.
Development of antibiotic resistance is a major concern for the continued use in fuel ethanol production. With so few antibiotics approved for use, development of multi-drug resistant bacterial strains would be extremely detrimental. Combination products are applying two of the available antibiotics at the same time increasing the risk of multi-drug resistance.5 In fact, research has shown that synergistic drug combinations increase the rate of resistance compared to the individual drugs by selecting for multi-drug resistant cells.6,7 However, alternating or cycling different antibiotics slows resistance development.8
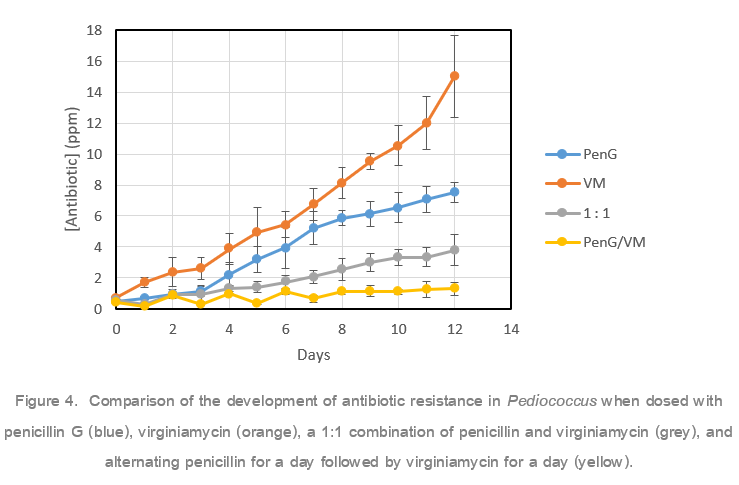
To further investigate how the antibiotic application method influences development of resistance, Pediococcus pentosaceus was once again grown in the presence of either penicillin, virginiamycin, a 1:1 pencillin to virginiamycin combination, or an alternating schedule of penicillin followed by virginiamycin. Pediococcus was grown in a series of wells with a gradient of antibiotic. Each day, cells from the highest concentration of antibiotic that still saw growth were used to inoculate a fresh set of media supplemented with an increased gradient of antibiotics (Figure 4). After a single day of growth, all antibiotic treatment methods tested showed growth at ≤ 0.7 ppm. After twelve days, both penicillin and virginiamycin exposed cells showed significant gain of resistance, 7.5 ppm and 15 ppm respectively. The 1:1 penicillin/virginiamycin blend exposed cells developed resistance to 3.8 ppm antibiotic. Alternating penicillin and virginiamycin was the most successful at limiting resistance development; these cells were resistant to only 1.3 ppm antibiotic. Only the alternating method kept development of resistance at or below typical antibiotic doses used in fuel ethanol fermentation.
Overall, our results show that penicillin G and virginiamycin when applied simultaneously act antagonistically against both lab bacterial strains and field isolates. Antibiotic combination products are more expensive, generally less effective, and increase the likelihood of multi-drug resistance. In addition to thorough CIP, use of a single antibiotic at a time with periodic switching to an alternative antibiotic, particularly one with a different mode of action, provides the most effective bacterial control while limiting development of multi-drug resistance.
References
1. Yeh P et al. Nature Genetics 2006;489–94.
2. Cokol M et al. Molecular Systems Biology 2011;544.
3. Johansen HK et al. Journal of Antimicrobial Chemotherapy 2000;973–80.
4. Tallarida RJ. Perspectives in Pharmacology 2012;2–8.
5. Bischoff KM et al. Journal of Industrial Microbiology and Biotechnology 2007;739–44.
6. Bollenbach T. Current Opinion in Microbiology 2015;1-9.
7. Pena-Miller R et al. PLOS Biology 2013;e1001540.
8. Kim S et al. PNAS 2014;14494–9.

